Resonance Assignment/Sparky: Difference between revisions
No edit summary |
No edit summary |
||
| Line 85: | Line 85: | ||
a1, a2, a3: add SW to peak in F1, F2, F3 dimension; for aliased spectra<br>A1, A2, A3: subtract SW from peak in F1, F2, F3 dimension; for aliased spectra<br>at: assignment tool; for assigning a peak<br>cl: adjust colour of ornament<br>ct: adjust contour levels and colours:<br>dr: delete resonances not used in any peak assignment (cleans up resonance list)<br>eu: undo last peak manipulation<br>it: integration tool<br>kr: restrictive peak picking tool<br>lt: opens the peak list for a given spectrum; various options in here<br>oc: ornament copy; for copying assignment/label information between spectra<br>ol: overlay views; useful for comparing spectra<br>op: ornament paste<br>oz: adjust size of ornament<br>pa: select all peaks in a spectrum<br>pc: peak center<br>pv: list of sizes and peak counts in all open spectra<br>rl: resonance list for the project; various functions and displays in there<br>rp: read in a list of peaks in sparky format (i.e. from AutoAssign or PINE)<br>rr: resonance rename<br>tb: table of resonances for the project: useful for seeing missing assignments<br>vc: view centering; useful for going to a specific peak in a 3D selected in a 2D<br>vd: view duplicate; duplicate view of a spectrum into another window<br>vR: show assignments on edge of spectrum<br>vS: show 1D slice on edge of spectrum<br>vt: view settings; a variety of spectral settings including aspect ratio<br>xa: show nucleus type on axis<br>xe: special Python command for saving peak list in xeasy format<br>xr: roll axes; useful for 3D and 4D spectra<br>xx: axis transpose<br>yt: synchronize axes of various spectra: useful for analyzing series of 3D’s<br>zf, zi, zo, zp: zoom full spectrum, zoom in, zoom out, zoom previous<br><br>Commonly used pointer modes (for the mouse):<br> F1: selection mode<br> F6: add a label<br> F7: add a line<br> F8: peak picking mode<br> F10: integration mode<br> F11: zoom mode | a1, a2, a3: add SW to peak in F1, F2, F3 dimension; for aliased spectra<br>A1, A2, A3: subtract SW from peak in F1, F2, F3 dimension; for aliased spectra<br>at: assignment tool; for assigning a peak<br>cl: adjust colour of ornament<br>ct: adjust contour levels and colours:<br>dr: delete resonances not used in any peak assignment (cleans up resonance list)<br>eu: undo last peak manipulation<br>it: integration tool<br>kr: restrictive peak picking tool<br>lt: opens the peak list for a given spectrum; various options in here<br>oc: ornament copy; for copying assignment/label information between spectra<br>ol: overlay views; useful for comparing spectra<br>op: ornament paste<br>oz: adjust size of ornament<br>pa: select all peaks in a spectrum<br>pc: peak center<br>pv: list of sizes and peak counts in all open spectra<br>rl: resonance list for the project; various functions and displays in there<br>rp: read in a list of peaks in sparky format (i.e. from AutoAssign or PINE)<br>rr: resonance rename<br>tb: table of resonances for the project: useful for seeing missing assignments<br>vc: view centering; useful for going to a specific peak in a 3D selected in a 2D<br>vd: view duplicate; duplicate view of a spectrum into another window<br>vR: show assignments on edge of spectrum<br>vS: show 1D slice on edge of spectrum<br>vt: view settings; a variety of spectral settings including aspect ratio<br>xa: show nucleus type on axis<br>xe: special Python command for saving peak list in xeasy format<br>xr: roll axes; useful for 3D and 4D spectra<br>xx: axis transpose<br>yt: synchronize axes of various spectra: useful for analyzing series of 3D’s<br>zf, zi, zo, zp: zoom full spectrum, zoom in, zoom out, zoom previous<br><br>Commonly used pointer modes (for the mouse):<br> F1: selection mode<br> F6: add a label<br> F7: add a line<br> F8: peak picking mode<br> F10: integration mode<br> F11: zoom mode | ||
== '''REFERENCES''' == | == '''REFERENCES''' == | ||
When using Sparky in your structure determination please cite:<br> T.D. Goddard & D.G. Kneller, SPARKY 3,<br> University of California, San Francisco<br><br> <br> <br> | When using Sparky in your structure determination please cite:<br> T.D. Goddard & D.G. Kneller, SPARKY 3,<br> University of California, San Francisco<br> | ||
JMA:10/27/09<br> <br> <br> | |||
<br> | <br> | ||
Revision as of 19:49, 27 October 2009
Sparky
INTRODUCTION
Sparky is a software package developed at the University of San Francisco for visualizing and analyzing multidimensional (2D, 3D or 4D) NMR spectra. Sparky plays a critical role as the link between spectral data and assignment/structure determination programs. The input for the program is processed spectra from various formats (NMRPipe, Felix, Varian or Bruker) converted into Sparky (i.e., “ucsf” format) via processing scripts supplied with the program. One can output a variety of data from Sparky including: chemical shift assignments, peak lists, and post script images of spectra. In the following sections we describe the practical aspects of using Sparky and standard approaches employed by NESG researchers using this program in structure determination projects. A descriptive on-line manual for a recent Sparky version (3.115) is available at: www.cgl.ucsf.edu/home/sparky/manual/index.html.
METHODS
1. Conversion of processed spectra to Sparky input format
Typically we process NMR spectra using NMRPipe, hence it is necessary to convert the spectra from pipe to ucsf format using the following command:
pipe2ucsf [options] filename.pipe filename.ucsf
For higher dimensional spectra a flag can be used to swap the dimension order if desired. For example “-213” swaps the order of the 1st and 2nd indirect dimensions of a 3D spectrum. This is applicable to many Varian 3D triple resonance spectra.
Another useful function is making 2D projections of higher dimensional spectra. This is done using the “ucsfdata” command. First, run the following command:
ucsfdata filename.ucsf
to obtain a list of information for each dimension of the spectrum. Then, for example, to make a 2D F23 projection of a 3D spectrum use:
ucsfdata –s1 [# points] –r –o output.ucsf input.ucsf
or
ucsfdata –p1 –o output.ucsf input.ucsf
2. Launching Sparky
To open a spectrum for the first time use:
sparky filename.ucsf
To open a single save file or project (a collection of save files) use:
sparky filename.save
sparky filename.proj
If using Sparky for the first time, a Sparky folder will be automatically generated in your home directory, with various subfolders. It is recommended that spectral (.ucsf), save and project files for a specific project be stored and launched from a specific directory in order to avoid confusion between multiple protein projects. The .save and .proj files are simple text files which contain links pointing to where data (.ucsf and .save files) are located on your machine. If a spectrum fails to open check the links in these files.
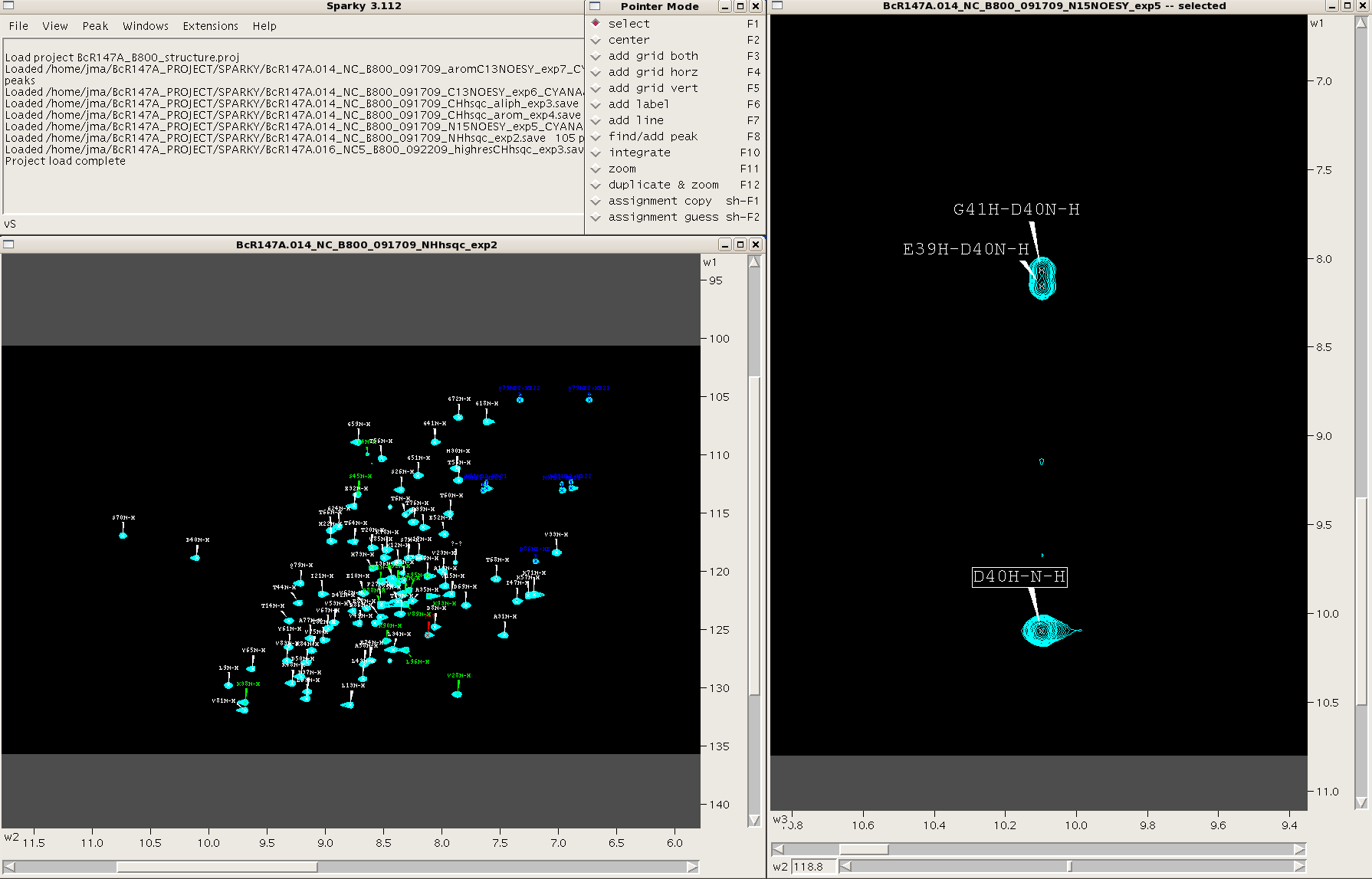
When Sparky is launched a menu box appears. This box tracks all commands performed during a session. One can make commands in Sparky using the mouse and clicking various commands under the different menus within the menu box or by using a plethora of two-letter “accelerators”. A “pointer mode” box also opens at the start of a session and allows the user to toggle between various pointer modes. Spectra are launched in individual windows. See Figure 1. All spectra are opened as 2D planes, with the additional dimension(s) of 3D and 4D spectra represented by sliders along the bottom of the spectral window.
Figure 1:
3. Peak Picking
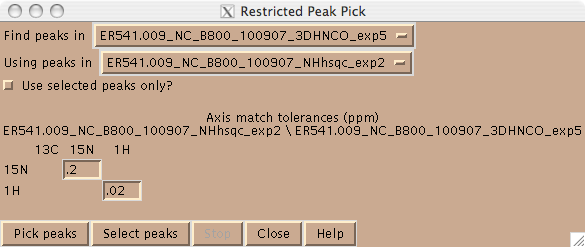
Peaks can be picked in Sparky in one of two ways. One can use the left mouse button in peak picking mode (F8). In this mode one can drag the mouse over the peak(s) in the spectrum and Sparky with place a peak at the maximum intensity. One can also place a peak using the left mouse button; this is desirable in cases where there is substantial peak overlap. Alternatively, one can pick peaks in a higher dimensional spectrum using peaks in a lower dimensional spectrum using the restrictive peak picking command “kr” and user defined tolerances (Figure 2). In this mode, be sure to set the contour levels in the target spectrum to an appropriate level using the “ct” command, in order to avoid picking a lot of noise.
Figure 2:
4. Manipulating and Analyzing Multiple Spectra
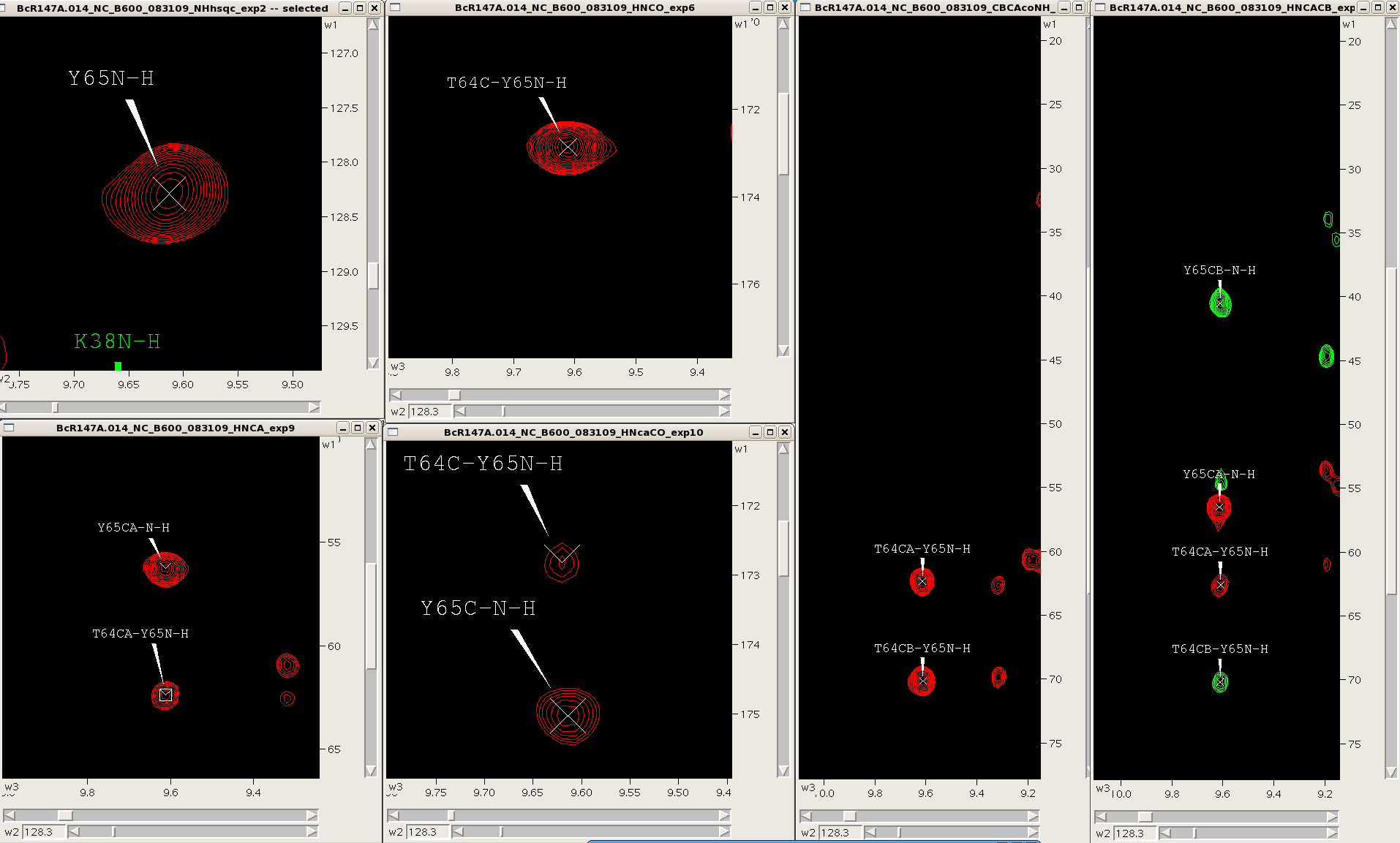
One can easily navigate through multiple spectra in Sparky, particularly spectra with common root dimensions (i.e., NH). For example, in the case of backbone resonance assignment with conventional triple resonance approaches one can synchronize 3D spectra (i.e., HNCO, HNcoCA, HNCA, HNcaCO, CBCAcoNH, HNCACB, HBHAcoNH) to each other and the NH-HSQC using the “yt” command, and then jump to successive NH-roots using the “vc” command (Figure 3). Similarly, one can use this approach to synchronize and navigate between entire spin systems in 3D side chain experiments (i.e., CCH-TOCSY, HCCH-TOCSY, HCCH-COSY) as well as NOESY spectra (i.e., 3D 15N-filtered and 13C-filtered NOESYs, and 4D 13C-13C-NOESY).
Figure 3:
5. Assignment and Peak Lists
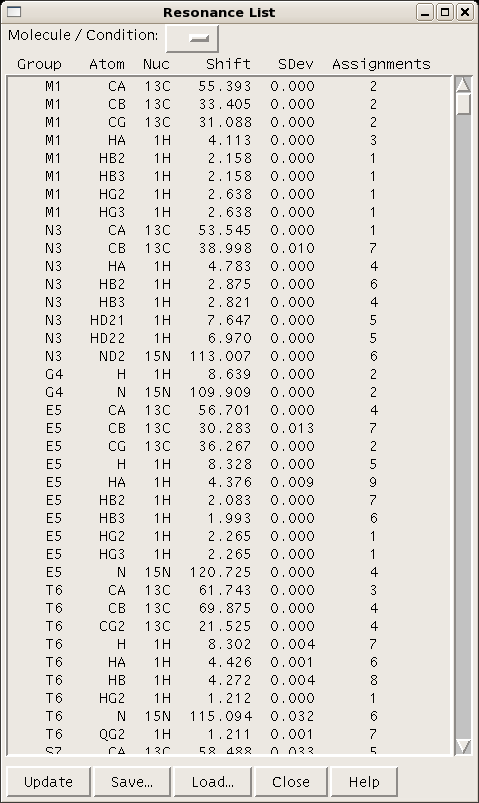
A table of resonance lists for a given project can be generated using the command “rl” (Figure 4).
Figure 4:
This list can be saved and subsequently converted to bmrb format using a script in the AutoAssign/bin directory.
sparkyRL2bmrb.pl input_rl output_bmrb sequence_start sequence [options]
options: -diastereo –unique_methyls –bmrb2
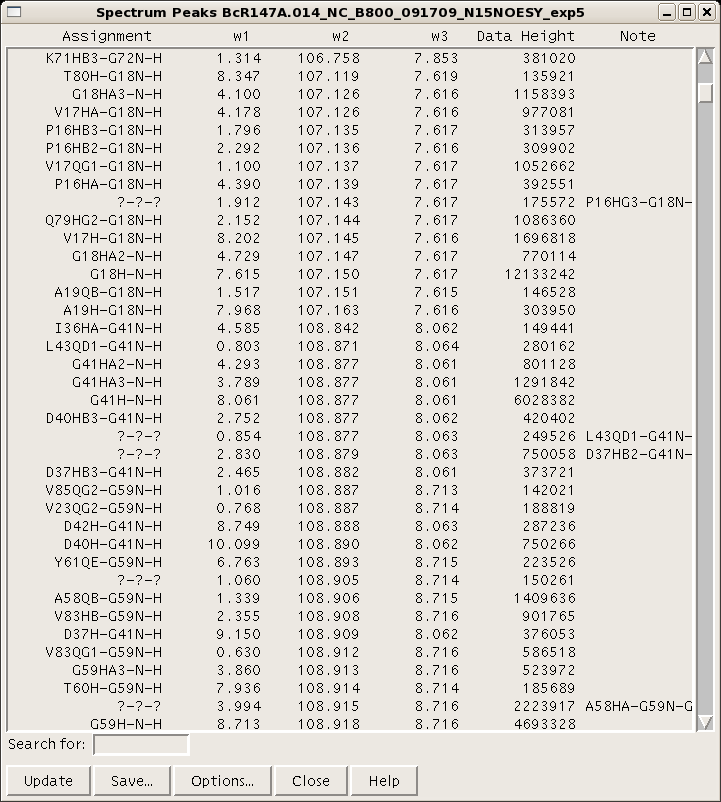
Each resonance value is an average of all the assignments for that resonance across all open spectra. Therefore, be sure that the assignment nomenclature used throughout the project is consistent. A peak list for a given spectrum can be generated and using the command “lt”. (Figure 5). There are various options for column visualization and ordering, and the list can be saved for input into AutoAssign or AutoStructure. For input into CYANA, NOESY spectral peak lists can be generated in XEASY format using the “xe” command.
Figure 5:
Assigned backbone peak lists from AutoAssign or PINE, and assigned NOESY peak lists from AutoStructure can be easily read back into the appropriate spectra using the “rp” command. In the case of CYANA peak lists one must convert the peak list back into Sparky using the following command:
awk –f peaks2sparky.awk file.prot file_cycle7.peaks > assigned_4sparky.list
The peaks2sparky.awk script can be obtained at:
www.nmr.chem.uu.nl/~eiso/help/index.html
6. Common Sparky Commands
There are numerous useful two-letter commands that can speed up spectral analysis in Sparky. Here is a list of commonly used accelerators:
a1, a2, a3: add SW to peak in F1, F2, F3 dimension; for aliased spectra
A1, A2, A3: subtract SW from peak in F1, F2, F3 dimension; for aliased spectra
at: assignment tool; for assigning a peak
cl: adjust colour of ornament
ct: adjust contour levels and colours:
dr: delete resonances not used in any peak assignment (cleans up resonance list)
eu: undo last peak manipulation
it: integration tool
kr: restrictive peak picking tool
lt: opens the peak list for a given spectrum; various options in here
oc: ornament copy; for copying assignment/label information between spectra
ol: overlay views; useful for comparing spectra
op: ornament paste
oz: adjust size of ornament
pa: select all peaks in a spectrum
pc: peak center
pv: list of sizes and peak counts in all open spectra
rl: resonance list for the project; various functions and displays in there
rp: read in a list of peaks in sparky format (i.e. from AutoAssign or PINE)
rr: resonance rename
tb: table of resonances for the project: useful for seeing missing assignments
vc: view centering; useful for going to a specific peak in a 3D selected in a 2D
vd: view duplicate; duplicate view of a spectrum into another window
vR: show assignments on edge of spectrum
vS: show 1D slice on edge of spectrum
vt: view settings; a variety of spectral settings including aspect ratio
xa: show nucleus type on axis
xe: special Python command for saving peak list in xeasy format
xr: roll axes; useful for 3D and 4D spectra
xx: axis transpose
yt: synchronize axes of various spectra: useful for analyzing series of 3D’s
zf, zi, zo, zp: zoom full spectrum, zoom in, zoom out, zoom previous
Commonly used pointer modes (for the mouse):
F1: selection mode
F6: add a label
F7: add a line
F8: peak picking mode
F10: integration mode
F11: zoom mode
REFERENCES
When using Sparky in your structure determination please cite:
T.D. Goddard & D.G. Kneller, SPARKY 3,
University of California, San Francisco
JMA:10/27/09