REDCRAFT
Introduction
Characterization of structure and dynamics of a biomolecule is crucial in better understanding of its function. It is difficult to separate structure from dynamics since the two are intimately related and any attempt in structure elucidation that disregards the dynamics (or vice versa) can produce faulty results. Newly reignited use of the RDC data has provided a potential avenue for characterizing structure and dynamics of proteins in biologically relevant timescales. This explosion in use of RDC data in structure determination of macromolecules is mainly due to its distinct advantages over the traditional distance constraints. In general, RDC data are more precise and easier to measure, while concurrently providing informative structural and motional information. Direct relationship between a carefully selected set of RDC data and structural parameters such as backbone torsion angles can be cited as one advantage of the RDC data. However, structure determination primarily based on RDC data requires new programs that operate in fundamentally different ways from those that use NOE data. REDCRAFT (REsidual Dipolar Coupling based Residue Assembly and Filtering Tool) has been developed for extraction and unraveling of the information reported by the RDC data [1]. Although REDCRAFT is capable of utilizing many advantages the RDC data, here we highlight only two distinct features.
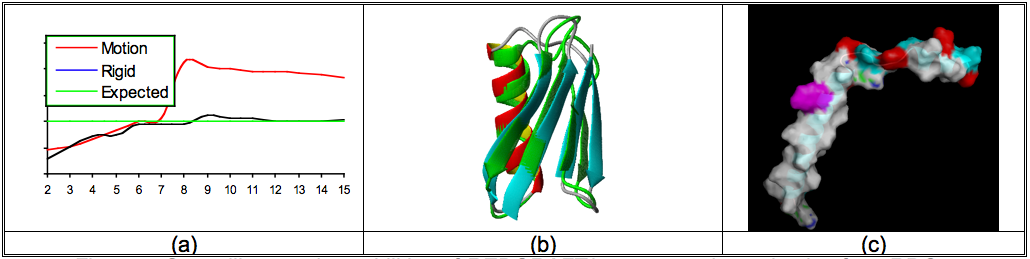
Simultaneous structure and Motion – Residual dipolar couplings encapsulate information regarding a protein's (in general any molecular system) anisotropic alignment, its structure, and internal motion/rigidity. REDCRAFT is capable of assembling the structure of a protein with the additional capability of identifying the location of internal motion in a theoretically well founded manner. In addition to an ensemble of protein structures, REDCRAFT produces a dynamic-profile that can be used to identify the location of internal motion as shown in Figure 1(a). Any sudden transition in this profile can be indicative of internal motion. Figure 1(a) displays the score of fitness to the experimental data (on the vertical axis) acquired from a Structural Genomics target protein as a function of the residue number (on the x-axis). The expected error (green line) is established based on the experimental noise. The blue line is the dynamic-profile for a fragment in the core of this protein that exhibits no internal motion while the red line is the dynamic-profile of the C-terminal fragment of the same protein; the presence of internal motion within the C-terminal region is clear. Identification of existence and location of internal motion has lead to separate structure determination of the C-terminal region (which is also an additional enabling feature of RECRAFT) of this protein. This separate treatment has produced a very well formed and dynamical helix that produces significantly reduced order parameters with respect to the core portion of this protein.
Figure 1: Some Applications of REDCRAFT in Protein Structure Determination
Structure determination of proteins from sparse data – RDC Data can be utilized as short-range and long-range constraints. This attribute of the RDC data can be used in order to reduce the overall data requirement for structure determination of macromolecules and more specifically challenging proteins for which only sparse amount of experimental data can be acquired. Although a reduction in the amount of required data for meaningful structure determination is generally welcomed, special class of proteins such as membrane, homo-multimeric or glycosylated proteins significantly benefit from this reduction. Membrane proteins are a challenging class of proteins because in general it is difficult to obtain the needed 12 or more NOEs per residue. On the other hand, collection of 2 or more RDCs per residue from multiple alignment media is within our reach. REDCRAFT has demonstrated its success in De Novo structure determination from only the backbone N-H and Cα-Hα RDCs in two or more alignment media. Figure 1(b) illustrates a superposition of the structure obtained from REDCRAFT (by using only N-H, Cα-Hα and C-N RDC data from two alignment media) and the deposited structure 1P7E by Bax's group. These two structures exhibit a 1.0 Å similarity over their backbone atoms. Furthermore, REDCRAFT is capable of accepting angular constraints such as TALOS constraints in order to further reduce the data requirement. The most recent application of REDCRAFT successfully produced the structure of a membrane associated form of the Pf1 coat protein Figure 1(c) by using the backbone N-H RDCs acquired in two alignment media and TALOS constraints.
Installation and Documentation
Descriptions for downloading, installing and using REDCRAFT can be found on-line.
References
[1 Bryson M, Tian F, Prestegard JH & Valafar H. Redcraft: a tool for simultaneous characterization of protein backbone structure and motion from rdc data. J. Magn. Reson. (2008) 191: pp. 322-334.]